Hydrophobicity
Hydrophobicity values are mapped according to the table with hydrophobicity values for 20 amino acids Radzicka et al. (1988) weighed by accessbility value of each residue, as follows:
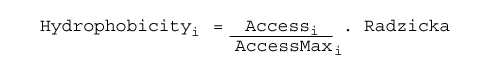
Where:
| Accessi | represents the accessbility for residue i |
| AccessMaxi | represents the the maximum accessibility value for residue i |
| Radzickai | is the Radzicka's hydrophobicity value for residue i. |
Placing the cursor above
this element: pop-up area will show the position (sequence number
and AA three letter code) for selected amino acid and the numerical
value for Hydrophobicity for this amino acid.
Left mouse click: no action
Right
mouse click: on any of the "Hydrophobicity" will generate following
menu and actions:
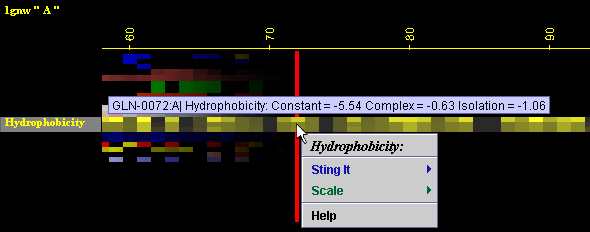
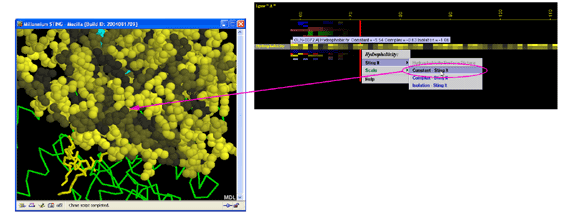
A) STING it :
This option generates structural presentation with color coding of the
amino acids corresponding to the "Hydrophobicity" color coding. Amino
acids are presented in CPK rendering.
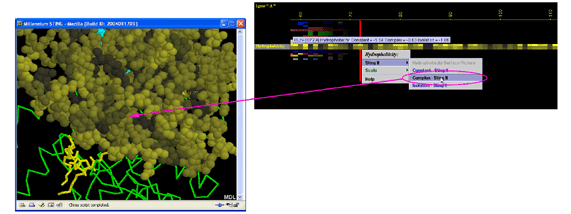
B) The Hydrophobicity
values shown at the picture below, were generated using the folowing table (Radzicka et al. (1988)).
Figure below shows the difference in concept when data display is in
question: GRASP does it one parameter at the time shown at the
molecular surface. BLUE STAR STING JPD does it all parameters at the same
time for all amino acids (including those at the surface).
| The hydrophobicity is very
important to consider when analyzing protein folding because it was
shown that the hydrophobic forces dominate the process.
Also, hydrophobic patches at the surface (see above HOT_SPOTS), are potentially important for identification of surface portions which can engage in the protein interactions/binding. |
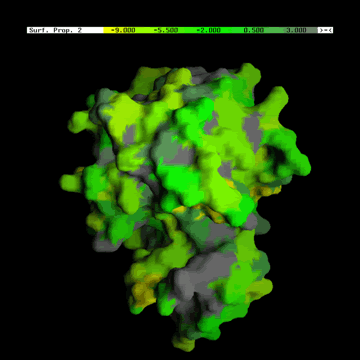 |
Table of hydropobicity values assigned to each amino acid
The table is taken from the work by Radzicka et al (1988)
|
Hydrophobicity |
Amino Acid
|
| -14.92 |
(Arg)
|
| -8.72 |
(Asp)yellow
|
| -6.81 |
(Glu)
|
| -6.64 |
(Asn)
|
| -5.55 |
(Lys)
|
| -5.54 |
(Gln)
|
| -4.66 |
(His)
|
| -3.40 |
(Ser)
|
| -2.57 |
(Thr)
|
| -0.14 |
(Tyr)
|
| 0.94 |
(Gly)
|
| 1.28 |
(Cys)
|
| 1.81 |
(Ala)
|
| 2.33 |
(Trp)
|
| 2.35 |
(Met)
|
| 2.98 |
(Phe)
|
| 3.50 |
(Pro)
|
| 4.04 |
(Val)
|
| 4.92 |
(Ile)
|
| 4.92 |
(Leu) gray
|